Cheaha GettingStarted deprecated: Difference between revisions
Jpr@uab.edu (talk | contribs) (→Hardware: Add ink to cluster schematic and improve description of resources) |
Jpr@uab.edu (talk | contribs) (→Access: Improve flow of configuration instructions) |
||
| Line 7: | Line 7: | ||
Usage of Cheaha is governed by [http://www.uabgrid.uab.edu/aup UAB's Acceptable Use Policy (AUP)] for computer resources. | Usage of Cheaha is governed by [http://www.uabgrid.uab.edu/aup UAB's Acceptable Use Policy (AUP)] for computer resources. | ||
The official DNS name of Cheaha's frontend machine is ''cheaha.uabgrid.uab.edu''. If you want to refer to the machine as ''cheaha'', you'll have to either add | The official DNS name of Cheaha's frontend machine is ''cheaha.uabgrid.uab.edu''. If you want to refer to the machine as ''cheaha'', you'll have to either add the "uabgrid.uab.edu" to you computer's DNS search path. On Unix-derived systems (Linux, Mac) you can edit your computers /etc/resolv.conf as follows (you'll need administrator access to edit this file) | ||
<pre> | <pre> | ||
search uabgrid.uab.edu | search uabgrid.uab.edu | ||
</pre> | </pre> | ||
to | Or you can customize your SSH configuration to use the short name "cheaha" as a connection name. On systems using OpenSSH you can add the following to your ~/.ssh/config file | ||
<pre> | <pre> | ||
| Line 17: | Line 17: | ||
Hostname cheaha.uabgrid.uab.edu | Hostname cheaha.uabgrid.uab.edu | ||
</pre> | </pre> | ||
== Hardware == | == Hardware == | ||
Revision as of 11:51, 16 September 2011
Information about the history and future plans for Cheaha is available on the Cheaha UABGrid Documentation page.
Access
To request an account on Cheaha, please submit an authorization request to the IT Research Computing staff. Please include some background information about the work you plan on doing on the cluster and the group you work with, ie. your lab or affiliation.
Usage of Cheaha is governed by UAB's Acceptable Use Policy (AUP) for computer resources.
The official DNS name of Cheaha's frontend machine is cheaha.uabgrid.uab.edu. If you want to refer to the machine as cheaha, you'll have to either add the "uabgrid.uab.edu" to you computer's DNS search path. On Unix-derived systems (Linux, Mac) you can edit your computers /etc/resolv.conf as follows (you'll need administrator access to edit this file)
search uabgrid.uab.edu
Or you can customize your SSH configuration to use the short name "cheaha" as a connection name. On systems using OpenSSH you can add the following to your ~/.ssh/config file
Host cheaha Hostname cheaha.uabgrid.uab.edu
Hardware
Cheaha a Dell PowerEdge 2950 with 16GB of RAM and two quad core Intel Xeon 3GHz processors. It provides an interactive Linux environment with access to cluster computing resources controlled by the SGE scheduler.
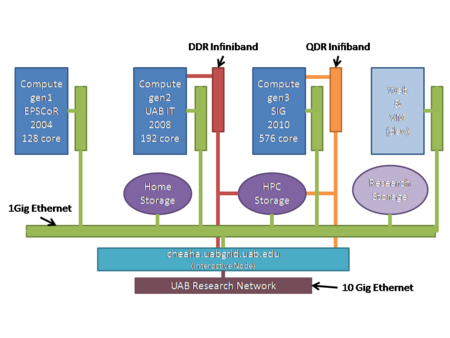
There are 896 cores available for batch computing. They are a collection of three generations of hardware:
- gen1: 128 1.8 GHz cores across 64 nodes each with 2 GB RAM and interconnected with 1Gbs Ethernet.
- gen2: 192 3.0GHz cores across 24 nodes each with 16GB RAM and interconnected with DDR Infiniband;
- gen3: 576 2.8 GHz cores across 48 nodes each with 48GB RAM and interconnected with QDR Infiniband;
Cheaha has a 240 TB Lustre high performance file system attached via Infiniband and GigE (depending on the networks available to the nodes, see hardware list above). The Lustre file system offers high performance access to large files in personal scratch and shared project directories. An additional 40TB are available for general research storage.
Further details on and a history of the hardware are available on the main Cheaha page.
Cluster Software
- Rocks 5.4
- CentOS 5.5 x86_64
- Grid Engine 6.2u5
- Globus 4.0.8
- Gridway 5.4.0
Storage
Home directories
Your home directory on Cheaha is NFS-mounted to the compute nodes as /home/$USER or $HOME. It is acceptable to use your home directory a location to store job scripts, custom code, libraries, job scripts.
The home directory must not be used to store large amounts of data.
Scratch
Research Computing policy requires that all bulky input and output must be located on the scratch space. The home directory is intended to store your job scripts, log files, libraries and other supporting files.
Cheaha has two types of scratch space, network mounted and local.
- Network scratch ($UABGRID_SCRATCH) is available on the head node and each compute node. This storage is a Lustre high performance file system providing roughly 240TB of storage. This should be your jobs primary working directory, unless the job would benefit from local scratch (see below).
- Local scratch is physically located on each compute node and is not accessible to the other nodes (including the head node). This space is useful if the job performs a lot of file I/O. Most of the jobs that run on our clusters do not fall into this category. Because the local scratch is inaccessible outside the job, it is important to note that you must move any data between local scratch to your network accessible scratch within your job. For example, step 1 in the job could be to copy the input from $UABGRID_SCRATCH to /scratch/$USER, step 2 code execution, step 3 move the data back to $UABGRID_SCRATCH.
Important Information:
- Scratch space (network and local) is not backed up.
- Research Computing expects each user to keep their scratch areas clean. The clusters are not to be used for archiving data.
Network Scratch
Network scratch is available using the environment variable $UABGRID_SCRATCH or directly by /lustre/scratch/$USER
It is advisable to use the environment variable whenever possible rather than the hard coded path.
Local Scratch
Local scratch is available on each compute node under /scratch.
Each compute node has a local scratch directory, /scratch. If your job performs a lot of file I/O, the job may run quicker (and possibly more stable) by using /scratch/$USER rather than reading and writing using your network mounted scratch directory. The amount of scratch space available on each compute node is approximately 40GB.
The following is a typical sequence of events within a job script using local scratch:
- Create a directory for your user called /scratch/$USER
mkdir -p /scratch/$USER/$JOB_ID
- Copy the data from $UABGRID_SCRATCH to /scratch/$USER
cp -a $UABGRID_SCRATCH/GeneData /scratch/$USER/$JOB_ID/
- Run the application
geneapp -S 1 -D 10 \< /scratch/$USER/$JOB_ID/GeneData \> /scratch/$USER/$JOB_ID/geneapp.out
- Delete anything that you don't want to move back to network scratch (for example the copy of the input data)
rm -rf /scratch/$USER/$JOB_ID/GeneData
- Move the data that you want to keep from local to network scratch
mv /scratch/$USER/$JOB_ID $UABGRID_SCRATCH/
The following is an example of what the code might look like in a job script (remember if the job is an array job, you may need to use /scratch/$USER/$JOB_ID/$SGE_TASK_ID to prevent multiple tasks from overwriting each other):
#!/bin/bash #$ -S /bin/bash #$ -cwd # #$ -N scratch_example #$ -pe smp 1 #$ -l h_rt=00:20:00,s_rt=00:18:00,vf=2G #$ -j y # #$ -M YOUR_EMAIL_ADDRESS #$ -m eas # module load R/R-2.9.0 if [ ! -d /scratch/$USER/$JOB_ID ]; then mkdir -p /scratch/$USER/$JOB_ID chmod 700 /scratch/$USER fi cp -a $UABGRID_SCRATCH/GeneData /scratch/$USER/$JOB_ID/ $HOME/bin/geneapp -S 1 -D 10 < /scratch/$USER/$JOB_ID/GeneData > /scratch/$USER/$JOB_ID/geneapp.out rm -rf /scratch/$USER/$JOB_ID/GeneData mv /scratch/$USER/$JOB_ID $UABGRID_SCRATCH/
By default, the $USER environment variable contains your login ID and the grid engine will populate $JOB_ID and $SGE_TASK_ID variables with the correct job and task IDs.
The mkdir command creates the full directory path (the -p switch is important). The chmod ensures that other users are not able to view files under this directory.
Please make sure to clean up the scratch space. This space is not to be used as a long term storage device for data and is subject to being erased without notice if the file systems fill up.
Project Storage
Cheaha has a location where shared data can be stored called $UABGRID_PROJECT
This is helpful if a team of researchers must access the same data. Please open a help desk ticket to request a project directory under $UABGRID_PROJECT.
Uploading Data
Data can be moved onto the cluster (pushed) from a remote client (ie. you desktop) via SCP or SFTP. Data can also be downloaded to the cluster (pulled) by issuing transfer commands once you are logged into the cluster. Common transfer methods are `wget <URL>`, FTP, or SCP, and depend on how the data is made available from the data provider.
Large data sets should be staged directly to your $UABGRID_SCRATCH directory so as not to fill up $HOME. If you are working on a data set shared with multiple users, it's preferable to request space in $UABGRID_PROJECT rather than duplicating the data for each user.
Environment Modules
Environment Modules is installed on Cheaha and should be used when constructing your job scripts if an applicable module file exists. Using the module command you can easily configure your environment for specific software packages without having to know the specific environment variables and values to set. Modules allows you to dynamically configure your environment without having to logout / login for the changes to take affect.
If you find that specific software does not have a module, please submit a helpdesk ticket to request the module.
Note: If you are using LAM MPI for parallel jobs, you must load the LAM module in both your job script and your profile. For example, assume we want to use LAM-MPI compiled for GNU:
- for BASH users add this to your ~/.bashrc and your job script, or for CSH users add this to your ~/.cshrc and your job script
module load lammpi/lam-7.1-gnu
- Cheaha supports bash completion for the module command. For example, type 'module' and press the TAB key twice to see a list of options:
module TAB TAB add display initlist keyword refresh switch use apropos help initprepend list rm unload whatis avail initadd initrm load show unuse clear initclear initswitch purge swap update
- To see the list of available modulefiles on the cluster, run the module avail command (note the example list below may not be complete!) or module load followed by two tab key presses:
module avail R/R-2.11.1 cufflinks/cufflinks-0.9 intel/intel-compilers mvapich-intel rna_pipeline/rna_pipeline-0.31 R/R-2.6.2 eigenstrat/eigenstrat jags/jags-1.0-gnu mvapich2-gnu rna_pipeline/rna_pipeline-0.5.0 R/R-2.7.2 eigenstrat/eigenstrat-2.0 lammpi/lam-7.1-gnu namd/namd-2.6 s.a.g.e./sage-6.0 R/R-2.8.1 ent/ent-1.0.2 lammpi/lam-7.1-intel namd/namd-2.7 samtools/samtools R/R-2.9.0 fastphase/fastphase-1.4 mach/mach openmpi/openmpi-1.2-gnu samtools/samtools-0.1 R/R-2.9.2 fftw/fftw3-gnu macs/macs openmpi/openmpi-1.2-intel shrimp/shrimp-1.2 RAxML/RAxML-7.2.6 fftw/fftw3-intel macs/macs-1.3.6 openmpi/openmpi-gnu shrimp/shrimp-1.3 VEGAS/VEGAS-0.8 freesurfer/freesurfer-4.5 maq/maq-0.7 openmpi/openmpi-intel spparks/spparks amber/amber-10.0-intel fregene/fregene-2008 marthlab/gigabayes openmpi-gnu structure/structure-2.2 amber/amber-11-intel fsl/fsl-4.1.6 marthlab/mosaik openmpi-intel tau/tau apbs/apbs-1.0 genn/genn marthlab/pyrobayes paraview/paraview-3.4 tau/tau-2.18.2p2 atlas/atlas gromacs/gromacs-4-gnu mathworks/R2009a paraview/paraview-3.6 tau/tau-lam-intel birdsuite/birdsuite-1.5.3 gromacs/gromacs-4-intel mathworks/R2009b pdt/pdt tophat/tophat birdsuite/birdsuite-1.5.5 hapgen/hapgen mathworks/R2010a pdt/pdt-3.14 tophat/tophat-1.0.8 bowtie/bowtie hapgen/hapgen-1.3.0 mpich/mpich-1.2-gnu phase/phase tophat/tophat-1.1 bowtie/bowtie-0.10 haskell/ghc mpich/mpich-1.2-intel plink/plink vmd/vmd bowtie/bowtie-0.12 illuminus/illuminus mpich/mpich2-gnu plink/plink-1.05 vmd/vmd-1.8.6 bowtie/bowtie-0.9 impute/impute mrbayes/mrbayes-gnu plink/plink-1.06 chase impute/impute-2.0.3 mrbayes/mrbayes-intel plink/plink-1.07 cufflinks/cufflinks impute/impute-2.1.0 mvapich-gnu python/python-2.6
Some software packages have multiple module files, for example:
- plink/plink
- plink/plink-1.05
- plink/plink-1.06
In this case, the plink/plink module will always load the latest version, so loading this module is equivalent to loading plink/plink-1.06. If you always want to use the latest version, use this approach. If you want use a specific version, use the module file containing the appropriate version number.
Some modules, when loaded, will actually load other modules. For example, the gromacs/gromacs-4-intel module will also load openmpi/openmpi-intel and fftw/fftw3-intel.
- To load a module, ex: for a Gromacs job, use the following module load command in your job script:
module load gromacs/gromacs-4-intel
- To see a list of the modules that you currently have loaded use the module list command
module list Currently Loaded Modulefiles: 1) fftw/fftw3-intel 2) openmpi/openmpi-intel 3) gromacs/gromacs-4-intel
- A module can be removed from your environment by using the module unload command:
module unload gromacs/gromacs-4-intel module list No Modulefiles Currently Loaded.
- The definition of a module can also be viewed using the module show command, revealing what a specific module will do to your environment:
module show gromacs/gromacs-4-intel ------------------------------------------------------------------- /etc/modulefiles/gromacs/gromacs-4-intel: module-whatis Sets up gromacs-intel v4.0.2 in your enviornment module load fftw/fftw3-intel module load openmpi/openmpi-intel prepend-path PATH /opt/uabeng/gromacs/intel/4/bin/ prepend-path LD_LIBRARY_PATH /opt/uabeng/gromacs/intel/4/lib prepend-path MANPATH /opt/uabeng/gromacs/intel/4/man -------------------------------------------------------------------
Sample Job Scripts
The following are sample job scripts, please be careful to edit these for your environment (i.e. replace YOUR_EMAIL_ADDRESS with your real email address), set the h_rt to an appropriate runtime limit and modify the job name and any other parameters.
Hello World
Hello World is the classic example used throughout programming. We don't want to buck the system, so we'll use it as well to demonstrate jobs submission.
Hello World (serial)
A serial job is one that can run independently of other commands, ie. it doesn't depend on the data from other jobs running simultaneously. You can run many serial jobs in any order. This is a common solution to processing lots of data when each command works on a single piece of data. For example, running the same conversion on 100's of images.
Here we show how to create job script for one simple command. Running more than one command just requires submitting more jobs.
- Create your hello world application. This will be a simple shell script that prints "Hello world!".
- Run this command to create a script (just copy and past it on to the command line).
cat > helloworld.sh << EOF echo "Hello World!" EOF chmod +x helloworld.sh
- Create the Grid Engine job script that will request 1 cpu slots and a maximum runtime of 10 minutes
$ vi helloworld.qsub
#!/bin/bash # # Define the shell used by your compute job # #$ -S /bin/bash # # Tell the cluster to run in the current directory from where you submit the job # #$ -cwd # # Name your job to make it easier for you to track # #$ -N HelloWorld_serial # # Tell the scheduler only need 10 minutes # #$ -l h_rt=00:10:00,s_rt=0:08:00 # # Set your email address and request notification when you job is complete or if it fails # #$ -M YOUR_EMAIL_ADDRESS #$ -m eas # # # Load the appropriate module files # # (no module is needed for this example, normally an appropriate module load command appears here) # # Tell the scheduler to use the environment from your current shell # #$ -V ./helloworld.sh
- Submit the job to Grid Engine and check the status using qstat
$ qsub helloworld.qsub
Your job 11613 ("HelloWorld") has been submitted
$ qstat -u $USER
job-ID prior name user state submit/start at queue slots ja-task-ID
-----------------------------------------------------------------------------------------------------------------
11613 8.79717 HelloWorld jsmith r 03/13/2009 09:24:35 all.q@compute-0-3.local 1
- When the job completes, you should have output files named HelloWorld.o* and HelloWorld.e* (replace the asterisk with the job ID, example HelloWorld.o11613). The .o file is the standard output from the job and .e will contain any errors.
$ cat HelloWorld.o11613 Hello World!
Hello World (parallel)
Example of above but using GNU Parallel command (coming soon).
Hello World (parallel with MPI)
MPI is used to coordinate the activity of many computations occurring in parallel. It is commonly used in simulation software for molecular dynamics, fluid dynamics, and similar domains where there is significant communication (data) exchanged between cooperating process.
Here is a simple parallel Grid Engine job script for running commands the rely on MPI. This example also includes the example of compiling the code and submitting the job script to the Grid Engine.
- First, create a directory for the Hello World jobs
$ mkdir -p ~/jobs/helloworld $ cd ~/jobs/helloworld
- Create the Hello World code written in C (this example of MPI enabled Hello World includes a 3 minute sleep to ensure the job runs for several minutes, a normal hello world example would run in a matter of seconds).
$ vi helloworld-mpi.c
#include <stdio.h>
#include <mpi.h>
main(int argc, char **argv)
{
int node;
int i, j;
float f;
MPI_Init(&argc,&argv);
MPI_Comm_rank(MPI_COMM_WORLD, &node);
printf("Hello World from Node %d.\n", node);
sleep(180);
for (j=0; j<=100000; j++)
for(i=0; i<=100000; i++)
f=i*2.718281828*i+i+i*3.141592654;
MPI_Finalize();
}
- Compile the code, first purging any modules you may have loaded followed by loading the module for OpenMPI GNU. The mpicc command will compile the code and produce a binary named helloworld_gnu_openmpi
$ module purge $ module load openmpi/openmpi-gnu $ mpicc helloworld-mpi.c -o helloworld_gnu_openmpi
- Create the Grid Engine job script that will request 8 cpu slots and a maximum runtime of 10 minutes
$ vi helloworld.qsub
#$ -S /bin/bash #$ -cwd # #$ -N HelloWorld #$ -pe openmpi 8 #$ -l h_rt=00:10:00,s_rt=0:08:00 #$ -j y # #$ -M YOUR_EMAIL_ADDRESS #$ -m eas # # Load the appropriate module files module load openmpi/openmpi-gnu #$ -V mpirun -np $NSLOTS helloworld_gnu_openmpi
- Submit the job to Grid Engine and check the status using qstat
$ qsub helloworld.qsub
Your job 11613 ("HelloWorld") has been submitted
$ qstat -u $USER
job-ID prior name user state submit/start at queue slots ja-task-ID
-----------------------------------------------------------------------------------------------------------------
11613 8.79717 HelloWorld jsmith r 03/13/2009 09:24:35 all.q@compute-0-3.local 8
- When the job completes, you should have output files named HelloWorld.o* and HelloWorld.po* (replace the asterisk with the job ID, example HelloWorld.o11613). The .o file is the standard output from the job and .po will contain any errors.
$ cat HelloWorld.o11613 Hello world! I'm 0 of 8 on compute-0-3.local Hello world! I'm 1 of 8 on compute-0-3.local Hello world! I'm 4 of 8 on compute-0-3.local Hello world! I'm 6 of 8 on compute-0-6.local Hello world! I'm 5 of 8 on compute-0-3.local Hello world! I'm 7 of 8 on compute-0-6.local Hello world! I'm 2 of 8 on compute-0-3.local Hello world! I'm 3 of 8 on compute-0-3.local
Gromacs
#!/bin/bash
#$ -S /bin/bash
#
# Request the maximum runtime for the job
#$ -l h_rt=2:00:00,s_rt=1:55:00
#
# Request the maximum memory needed for each slot / processor core
#$ -l vf=256M
#
# Send mail only when the job ends
#$ -m e
#
# Execute from the current working directory
#$ -cwd
#
#$ -j y
#
# Job Name and email
#$ -N G-4CPU-intel
#$ -M YOUR_EMAIL_ADDRESS
#
# Use OpenMPI parallel environment and 4 slots
#$ -pe openmpi 4
#
# Load the appropriate module(s)
. /etc/profile.d/modules.sh
module load gromacs/gromacs-4-intel
#
#$ -V
#
# Single precision
MDRUN=mdrun_mpi
# The $NSLOTS variable is set automatically by SGE to match the number of
# slots requests
export MYFILE=production-Npt-323K_${NSLOTS}CPU
cd ~/jobs/gromacs
mpirun -np $NSLOTS $MDRUN -v -np $NSLOTS -s $MYFILE -o $MYFILE -c $MYFILE -x $MYFILE -e $MYFILE -g ${MYFILE}.log
R
If you are using LAM MPI for parallel jobs, you must add the following two lines to your ~/.bashrc or ~/.cshrc file.
module load lammpi/lam-7.1-gnu
The following is an example job script that will use an array of 1000 tasks (-t 1-1000), each task has a max runtime of 2 hours and will use no more than 256 MB of RAM per task (h_rt=2:00:00,vf=256M)
The array is also throttled to only run 32 concurrent tasks at any time (-tc 32), this feature is not available on coosa.
More R examples are available here: Running R Jobs on a Rocks Cluster
Create a working directory and the job submission script
$ mkdir -p ~/jobs/ArrayExample $ cd ~/jobs/ArrayExample $ vi R-example-array-job.qsub
#!/bin/bash #$ -S /bin/bash # # Request the maximum runtime for the job #$ -l h_rt=2:00:00,s_rt=1:55:00 # # Request the maximum memory needed for each slot / processor core #$ -l vf=256M # #$ -M YOUR_EMAIL_ADDRESS # Email me only when tasks abort, use '#$ -m n' to disable all email for this job #$ -m a #$ -cwd #$ -j y # # Job Name #$ -N ArrayExample # #$ -t 1-1000 #$ -tc 32 # #$ -e $HOME/negcon/rep$TASK_ID/$JOB_NAME.e$JOB_ID.$TASK_ID #$ -o $HOME/negcon/rep$TASK_ID/$JOB_NAME.o$JOB_ID.$TASK_ID . /etc/profile.d/modules.sh module load R/R-2.9.0 #$ -v PATH,R_HOME,R_LIBS,LD_LIBRARY_PATH,CWD cd ~/jobs/ArrayExample/rep$SGE_TASK_ID R CMD BATCH rscript.R
Submit the job to the Grid Engine and check the status of the job using the qstat command
$ qsub R-example-array-job.qsub $ qstat
Installed Software
A partial list of installed software with additional instructions for their use is available on the Cheaha Software page.