Cheaha
https://docs.rc.uab.edu/
Please use the new documentation url https://docs.rc.uab.edu/ for all Research Computing documentation needs.
As a result of this move, we have deprecated use of this wiki for documentation. We are providing read-only access to the content to facilitate migration of bookmarks and to serve as an historical record. All content updates should be made at the new documentation site. The original wiki will not receive further updates.
Thank you,
The Research Computing Team
|
HPC Web Portal now in Beta The new HPC web portal is now available. We encourage you to try it out as an alternative to traditional clients. It provides file, shell, and desktop access to the cluster within your web browser. |
Cheaha is a campus resource dedicated to enhancing research computing productivity at UAB. Cheaha is managed by UAB Information Technology's Research Computing group (UAB ITRC) and is available to members of the UAB community in need of increased computational capacity. Cheaha supports high-performance computing (HPC) and high throughput computing (HTC) paradigms.
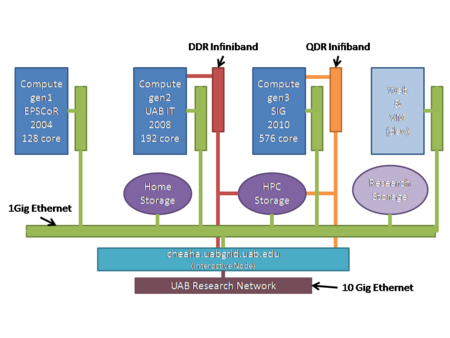
Cheaha provides users with a traditional command-line interactive environment with access to many scientific tools that can leverage its dedicated pool of local compute resources. Alternately, users of graphical applications can start a cluster desktop. The local compute pool provides access to three generations of compute hardware based on the x86-64 64-bit architecture. The compute resources are organized into a unified Research Computing System. The compute fabric for this system is anchored by the Cheaha cluster, a commodity cluster with several generations of hardware with approximately 3000 cores connected by low-latency Fourteen Data Rate (FDR) and Quad Data Rate (QDR) InfiniBand networks. The compute nodes are backed by 6.6PB raw GPFS storage on DDN SFA12KX hardware, a 180TB of high performance Lustre storage on a DDN hardware and an additional 20TB available for home directories on a traditional Hitachi SAN and other ancillary services. The compute nodes combine to provide over 120TFlops of dedicated computing power.
Cheaha is composed of resources that span data centers located in the UAB Shared Computing facility UAB 936 Building and the RUST Computer Center. Resource design and development is lead by UAB IT Research Computing in open collaboration with community members. Operational support is provided by UAB IT's Research Computing group.
Cheaha is named in honor of Cheaha Mountain, the highest peak in the state of Alabama. Cheaha is a popular destination whose summit offers clear vistas of the surrounding landscape. (Cheaha Mountain photo-streams on Flikr and Picasa).
Using
Getting Started
For information on getting an account, logging in, and running a job, please see Getting Started.
History
2005
In 2002 UAB was awarded an infrastructure development grant through the NSF EPsCoR program. This led to the 2005 acquisition of a 64 node compute cluster with two AMD Opteron 242 1.6Ghz CPUs per node (128 total cores). This cluster was named Cheaha. Cheaha expanded the compute capacity available at UAB and was the first general-access resource for the community. It lead to expanded roles for UAB IT in research computing support through the development of the UAB Shared HPC Facility in BEC and provided further engagement in Globus-based grid computing resource development on campus via UABgrid and regionally via SURAgrid.
2008
In 2008, money was allocated by UAB IT for hardware upgrades which lead to the acquisition of an additional 192 cores based on a Dell clustering solution with Intel Quad-Core E5450 3.0Ghz CPU in August of 2008. This uprade migrated Cheaha's core infrastructure to the Dell blade clustering solution. It provided a 3 fold increase in processor density over the original hardware and enables more computing power to be located in the same physical space with room for expansion, an important consideration in light of the continued growth in processing demand. This hardware represented a major technology upgrade that included space for additional expansion to address over-all capacity demand and enable resource reservation.
The 2008 upgrade began a continuous resource improvement plan that includes a phased development approach for Cheaha with on-going increases in capacity and feature enhancements being brought into production via an open community process.
Software improvements rolled into the 2008 upgrade included grid computing services to access distributed compute resources and orchestrate jobs using the GridWay meta-scheduler. An initial 10Gigabit Ethernet link establishing the UABgrid Research Network was designed to supports high speed data transfers between clusters connected to this network.
2009
In 2009, annual investment funds were directed toward establishing a fully connected dual data rate Infiniband network between the compute nodes added in 2008 and laying the foundation for a research storage system with a 60TB DDN storage system accessed via the Lustre distributed file system. The Infiniband and storage fabrics were designed to support significant increases in research data sets and their associate analytical demand.
2010
In 2010, UAB was awarded an NIH Small Instrumentation Grant (SIG) to further increase analytical and storage capacity. The grant funds were combined with the annual investment funds adding 576 cores (48 nodes) based on the Intel Westmere 2.66 GHz CPU, a quad data rate Infiniband fabric with 32 uplinks, an additional 120 TB of storage for the DDN fabric, and additional hardware to improve reliability. Additional improvements to the research compute platform involved extending the UAB Research Network to link the BEC and RUST data centers, adding 20TB of user and ancillary services storage
2012
In 2012, UAB IT Research Computing invested in the foundation hardware to expand long term storage and virtual machine capabilities with aqcuisition of 12 Dell 720xd system, each containing 16 cores, 96GB RAM, and 36TB of storage, creating a 192 core and 432TB virtual compute and storage fabric.
Additionaly hardware investment by the School of Public Health's Section on Statistical Genetics added three 384GB large memory nodes and an additional 48 cores to the QDR Infiniband fabric.
2013
In 2013, UAB IT Research Computing acquired an OpenStack cloud and Ceph storage software fabric through a partnership between Dell and Inktank in order to extend cloud computing solutions to the researchers at UAB and enhance the interfacing capabilities for HPC. This fabric is under active development and will see feature releases in the coming months.
2015/2016
In 2015/2016 UAB IT Research computing acquired 96 2x12 core (2304 cores total) 2.5 GHz Intel Xeon E5-2680 v3 compute nodes with FDR InfiniBand interconnect. Out of the 96 compute nodes, 36 nodes have 128 GB RAM, 38 nodes have 256 GB RAM, and 14 nodes have 384 GB RAM. There are also four compute nodes with the Intel Xeon Phi 7210 accelerator cards and four compute nodes with the NVIDIA K80 GPUs. More information can be found at Resources.
Grant and Publication Resources
The following description may prove useful in summarizing the services available via Cheaha. If you are using Cheaha for grant funded research please send information about your grant (funding source and grant number), a statement of intent for the research project and a list of the applications you are using to UAB IT Research Computing. If you are using Cheaha for exploratory research, please send a similar note on your research interest. Finally, any publications that rely on computations performed on Cheaha should include a statement acknowledging the use of UAB Research Computing facilities in your research, see the suggested example below. Please note, your acknowledgment may also need to include an addition statement acknowledging grant-funded hardware. We also ask that you send any references to publications based on your use of Cheaha compute resources.
Description of Cheaha for Grants
UAB IT Research Computing maintains high performance compute and storage resources for investigators. The Cheaha compute cluster provides 3120 conventional CPU cores across five generations of hardware that provide over 120 TFLOP/s of combined computational performance, and 20 TB of system memory interconnected via an Infiniband network. A high-performance, 6.6PB raw GPFS storage on DDN SFA12KX hardware and 180TB Lustre parallel file system built on a Direct Data Network (DDN) hardware platform is also connected to these cores via the Infiniband fabric. An additional 20TB of traditional SAN storage and 432TB of OpenStack+Ceph storage is available via a 10+ GigE network fabric. This general access compute fabric is available to all UAB investigators.
Additionally, NIH funded investigators are granted priority access to the NIH SIG award acquired compute and storage pool that includes an additional 576 2.66GHz Intel-based compute cores, 2.3TB RAM and 180TB in the high-performance Lustre parallel file system all interconnected via a QDR Infiniband network fabric.
Acknowledgment in Publications
This work was supported in part by the research computing resources acquired and managed by UAB IT Research Computing. Any opinions, findings and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the University of Alabama at Birmingham.
Additionally, NIH funded researchers using the cluster for genomics or proteomics analyses should include the following statement regarding the NIH funded gen3 (SIG) compute resources.
"Computational portions of this research were supported by NIH S10RR026723. Any opinions, findings and conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of the National Institutes of Health"
System Profile
Performance
| Generation | Type | Nodes | CPUs per Node | Cores Per CPU | Total Cores | Clock Speed (GHz) | Instructions Per Cycle | Hardware Reference |
|---|---|---|---|---|---|---|---|---|
| Gen 6† | Intel Xeon E5-2680 v3 | 96 | 2 | 12 | 2304 | 2.50 | 16 | Intel Xeon E5-2680 v3 |
| Gen 7†† | Intel Xeon E5-2680 v4 | 18 | 2 | 14 | 504 | 2.40 | 16 | Intel Xeon E5-2680 v4 |
| Gen 8 | Intel Xeon E5-2680 v4 | 35 | 2 | 12 | 840 | 2.50 | 16 | Intel Xeon E5-2680 v3 |
| Gen 9 | Intel Xeon Gold 6248R | 52 | 2 | 24 | 2496 | 3.0 | 16 | 3.0GHz Intel Xeon Gold 6248R |
| Generation | Theoretical Peak Tera-FLOPS |
|---|---|
| Gen 6† | 110 |
| Gen 7†† | 358 |
| Gen 8 | TBD |
| Gen 9 | TBD |
† Includes four Intel Xeon Phi 7210 accelerators and four NVIDIA K80 GPUs.
†† Includes 72 NVIDIA Tesla P100 16GB GPUs.
Hardware
The Cheaha Compute Platform includes three generations of commodity compute hardware, totaling 868 compute cores, 2.8TB of RAM, and over 200TB of storage.
The hardware is grouped into generations designated gen1, gen2, and gen3 (oldest to newest). The following descriptions highlight the hardware profile for each generation.
Generation 1 (gen1) -- 64 2-CPU AMD 1.6 GHz compute nodes with Gigabit interconnect. This is the original hardware collection purchased with NSF EPSCoR funds in 2005, approx $150K. These nodes are sometimes called the "Verari" nodes. These nodes are tagged as "verari-compute-#-#" in the ROCKS naming convention.(gen1 was decommissioned in June 2013)- Generation 2 (gen2) -- 24 2x4 core (196 cores total) Intel 3.0 GHz Intel compute nodes with dual data rate Infiniband interconnect and the initial high-perf storage implementation using 60TB DDN. This is the hardware collection purchased exclusively with the annual VPIT funds allocation, approx $150K/yr for the 2008 and 2009 fiscal years. These nodes are sometimes confusingly called "cheaha2" or "cheaha" nodes. These nodes are tagged as "cheaha-compute-#-#" in the ROCKS naming convention.
- Generation 3 (gen3) -- 48 2x6 core (576 cores total) 2.66 GHz Intel compute nodes with quad data rate Infiniband, ScaleMP, and the high-perf storage build-out for capacity and redundancy with 120TB DDN. This is the hardware collection purchased with a combination of the NIH SIG funds and some of the 2010 annual VPIT investment. These nodes were given the code name "sipsey" and tagged as such in the node naming for the queue system. These nodes are tagged as "sipsey-compute-#-#" in the ROCKS naming convention. 16 of the gen3 nodes (sipsey-compute-0-1 thru sipsey-compute-0-16) were upgraded in 2014 from 48GB to 96GB of memory per node.
- Generation 4 (gen4) -- 3 16 core (48 cores total) compute nodes. This hardware collection purchase by Hemant Tiwari of SSG. These nodes were given the code name "ssg" and tagged as such in the node naming for the queue system. These nodes are tagged as "ssg-compute-0-#" in the ROCKS naming convention.
- Generation 6 (gen6) --
- 26 Compute Nodes with two 12 core processors (Intel Xeon E5-2680 v3 2.5GHz) with 256GB DDR4 RAM, FDR InfiniBand and 10GigE network cards
- 14 Compute Nodes with two 12 core processors (Intel Xeon E5-2680 v3 2.5GHz) with 384GB DDR4 RAM, FDR InfiniBand and 10GigE network card
- FDR InfiniBand Switch
- 10Gigabit Ethernet Switch
- Management node and gigabit switch for cluster management
- Bright Advanced Cluster Management software licenses
Summarized, Cheaha's compute pool includes:
- gen4 is 48 cores of 2.70GHz eight-core Intel Xeon E5-2680 processors with 24G of RAM per core or 384GB total
- gen3.1 is 192 cores of 2.67GHz six-core Intel Xeon X5650 processors with 8Gb RAM per core or 96GB total
- gen3 is 384 cores of 2.67GHz six-core Intel Xeon X5650 processors with 4Gb RAM per core or 48GB total
- gen2 is 192 cores of 3.0GHz quad-core Intel Xeon E5450 processors with 2Gb RAM per core
gen1 is 100 cores of 1.6GhZ AMD Opteron 242 processors with 1Gb RAM per core(decomissioned June 2013)
| gen | queue | #nodes | cores/node | RAM/node |
|---|---|---|---|---|
| gen6.1 | ?? | 26 | 24 | 256G |
| gen6.2 | ?? | 14 | 24 | 384G |
| gen5 | openstack(?) | ? | ? | ?G |
| gen4 | ssg | 3 | 16 | 385G |
| gen3.1 | sipsey | 16 | 12 | 96G |
| gen3 | sipsey | 32 | 12 | 48G |
| gen2 | cheaha | 24 | 8 | 16G |
Software
Details of the software available on Cheaha can be found on the Cheaha cluster configuration page, an overview follows.
Cheaha uses Environment Modules to support account configuration. Please follow these specific steps for using environment modules.
Cheaha's software stack is built with the ROCKS, a Linux-based cluster distribution. Cheaha's operating system is CentOS with the following major cluster components:
A brief summary of the some of the available computational software and tools available includes:
- Amber
- FFTW
- Gromacs
- GSL
- NAMD
- VMD
- Intel Compilers
- GNU Compilers
- Java
- R
- OpenMPI
- MATLAB
Network
Cheaha is connected to the UAB Research Network which provides a dedicated 10Gbs networking backplane between clusters located the UAB Shared Computing Facility and Department of Computer and Information Science HPC Center. At present only Cheaha and Ferrum are connected via these 10Gbs interfaces. Data transfers rates of almost 8Gbs between these hosts have been demonstrated using Grid FTP, a multi-channel file transfer service that is used by GridWay to move data between clusters as part of the job management operations. This performance promises very efficient job management and the seamless integration of other clusters as connectivity to the research network is expanded.
Benchmarks
The continuous resource improvement process involves collecting benchmarks of the system. One of the measures of greatest interest to users of the system are benchmarks of specific application codes. The following benchmarks have been performed on the system and will be further expanded as additional benchmarks are performed.
Performance Statistics
Cheaha uses Ganglia to report cluster performance data. This information provides a helpful overview of the current and historical operating stats for Cheaha. You can access the Ganglia monitoring page here.
Availability
Cheaha is a general-purpose computer resource made available to the UAB community by UAB IT. As such, it is available for legitimate research and educational needs and is governed by UAB's Acceptable Use Policy (AUP) for computer resources.
Many software packages commonly used across UAB are available via Cheaha.
To request access to Cheaha, please send a request to send a request to the cluster support group.
Cheaha's intended use implies broad access to the community, however, no guarantees are made that specific computational resources will be available to all users. Availability guarantees can only be made for reserved resources.
Secure Shell Access
Please configure you client secure shell software to use the official host name to access Cheaha:
cheaha.uabgrid.uab.edu
To ensure that you are connecting to the legitimate host, you can verify that the fingerprint presented by your secure shell client for cheaha.uabgrid.uab.edu matches:
d4:2e:cc:12:95:a2:39:cc:b7:2c:d8:97:37:75:e9:6f
Scheduling Framework
Cheaha provides performance and management improvements for scientific workflows by enabling access to the processing power of multiple clusters through the use of the GridWay scheduling framework. GridWay enables the development of scientific workflows that leverage all computing resources available to the researcher and that can be controlled through a single management interface. This feature puts state-of-the-art technology in the hands of the research community.
Enhancements to Cheaha in general, and, in particular, its scheduling framework are intended to remain transparent to the user community. Cheaha is first and foremost a resource for predictable and dependable computation. In this spirit, Cheaha can continue to be viewed as a traditional HPC cluster that supports job management via Sun Grid Engine (SGE). User's who have no need for or interest in maximized access to computational resources can continue using the familiar SGE scheduling framework to manage compute jobs on Cheaha, with the familiar restriction that SGE managed jobs can only leverage the processing power of Cheaha's local compute pool. That is, these jobs, as in the past, cannot leverage cycles available on other clusters.
SGE
Cheaha provides access to its local compute pool via the SGE scheduler. This arrangement is identical to the existing HPC clusters on campus and mirrors the long-established configuration of Cheaha. Researchers experienced with other SGE-based clusters should find no difficulty leveraging this feature. For more information on getting started with SGE on Cheaha please see the Cheaha getting started page.
GridWay
Cheaha provides enhanced scientific workflow management and development capabilities via the GridWay scheduling framework. GridWay enables the orchestration of scientific workflows across multiple clusters. The pool of resources available as part in Phase 1 includes Cheaha, Olympus, Everest, and Ferrum. This pool can be monitored by any user of Cheaha by executing the gwhost command when logged into Cheaha.
The GridWay framework provides two interfaces. A scheduler interface similar to SGE is recommended for initial exploration and ordinary use. The scheduler activity can be monitored with the `gwps` command. Job submit and monitoring commands initiate and control commands described in a job description file. Outside of slightly different commands, the job description file operates as a template where specific fields are populated to affect the operation of the scheduler. These templates are less ambiguous than traditional SGE job scripts and can provide a direct migration path from SGE. A more subtle difference is that an explicit (though automated) job staging step is involved in order to start jobs. This can require more explicit handling of input and output files than is ordinarily required by SGE.
Additionally, very powerful programatic control is available via the DRMAA API. DRMMA enables the development of advanced scientific workflows that can leverage any number of computational resources. GridWay provides bindings to many popular programming languages like C, Java, Perl, Python and Ruby.
Both the traditional scheduler-based interface and the DRMMA API have been explored by development groups on-campus during the pilot evaluation phase of GridWay.
- UAB IT, UAB's School of Public Health Section on Statistical Genetics (SSG), and CIS established an interest group for R which leveraged the scheduler-based interface to improve the performance of select R language statistical analysis workflows.
- The CIS Collaborative Computing Laboratory has heavily leveraged the Java DRMMA API to develop DynamicBLAST, a scientific workflow to orchestrate and maximize the performance of BLAST across multiple resources.
GridWay Adoption
Adoption of GridWay is encouraged and future compute capacity enhancements will leverage the inherent flexibilities of GridWay. The nature of any new technology, however, implies a learning curve. The learning curve need not be steep and direct migration of basic SGE scripts is possible. Additionally, all Cheaha accounts are configured to support the use of GridWay as an alternative scheduler, empowering the adventurous.
Some important points worth considering in evaluating adoption of GridWay.
- GridWay cannot perform magic. If you ordinarily do not have access to other clusters or your code does not (or will not) run on a targeted cluster, GridWay cannot solve these problems for you. You must ensure your codes run on all compute resources you intend to include in your scheduling pool prior to submitting jobs to those resources.
- Migration of MPI jobs to a multi-cluster environment may involve additional effort. If you simply use MPI to coordinate the workers (rather than for low-latency peer communication), you should generally be able to structure your job to work across cluster boundaries. Otherwise, additional effort may be required to divide your data into smaller work units. It should be noted, however, that MPI itself cannot be used to communicate across cluster boundaries. Jobs distributed across cluster boundaries that leverage MPI internally must be sub-divided to run within isolated communication domains.
The UABgrid User Community stands ready to help you with GridWay adoption. If you are interested in exploring or adapting your workflows to use GridWay please subscribe to the UABgrid-User list and ask all questions there. Please do not submit GridWay-related questions via the ordinary cluster support channels. Additional on-line documentation will be developed to provide migration examples to help you further explore the power of GridWay.
Support
Operational support for Cheaha is provided by the Research Computing group in UAB IT. For questions regarding the operational status of Cheaha please send your request to support@vo.uabgrid.uab.edu. As a user of Cheaha you will automatically by subscribed to the hpc-announce email list. This subscription is mandatory for all users of Cheaha. It is our way of communicating important information regarding Cheaha to you. The traffic on this list is restricted to official communication and has a very low volume.
We have limited capacity, however, to support non-operational issue like "How do I write a job script" or "How do I compile a program". For such requests, you may find it more fruitful to send your questions to the hpc-users email list and request help from our peers in the HPC community at UAB. As with all mailing lists, please observe common mailing etiquette.
Finally, please remember that as you learned about HPC from others it becomes part of your responsibilty to help others on their quest. You should update this documentation or respond to mailing list requests of others.
You can subscribe to hpc-users by sending an email to:
sympa@vo.uabgrid.uab.edu with the subject subscribe hpc-users.
You can unsubribe from hpc-users by sending an email to:
sympa@vo.uabgrid.uab.edu with the subject unsubscribe hpc-users.
You can review archives of the list in the web hpc-archives.
If you need help using the list service please send an email to:
sympa@vo.uabgrid.uab.edu with the subject help
If you have questions about the operation of the list itself, please send an email to the owners of the list:
sympa@vo.uabgrid.uab.edu with a subject relavent to your issue with the list
If you are interested in contributing to the enhancement of HPC features at UAB or would like to talk to other cluster administrators, please join the hpc developers community at UAB.