Galaxy DNA-Seq Tutorial: Difference between revisions
Jump to navigation
Jump to search
No edit summary |
(FastQC and Compute stats cleanup) |
||
| Line 40: | Line 40: | ||
* The current list of genomes available at UAB is [http://docs.uabgrid.uab.edu/wiki/Galaxy#Available_datasets here] | * The current list of genomes available at UAB is [http://docs.uabgrid.uab.edu/wiki/Galaxy#Available_datasets here] | ||
* Used for a variety of tools, it can be overridden earlier | * Used for a variety of tools, it can be overridden earlier | ||
* Any problems with CMV-21950R_3_1.fastq ? | |||
== Assessing the quality of the data == | == Assessing the quality of the data == | ||
| Line 47: | Line 48: | ||
* Compute Quality Statistics | * Compute Quality Statistics | ||
FastQC gives an attractive visual output and will flag potential problems. | |||
=== FastQC === | |||
FastQC gives an attractive visual output and will flag potential problems. | |||
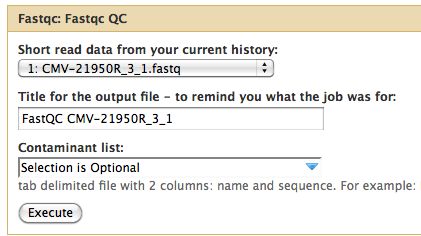
* Select NGS: QC and manipulation -> FastQC | |||
[[File:FastQCRunSettings.jpg]] | |||
* Run it on all the fastq files | |||
* Remember, you are on a cluster, this can be done in parrallel | |||
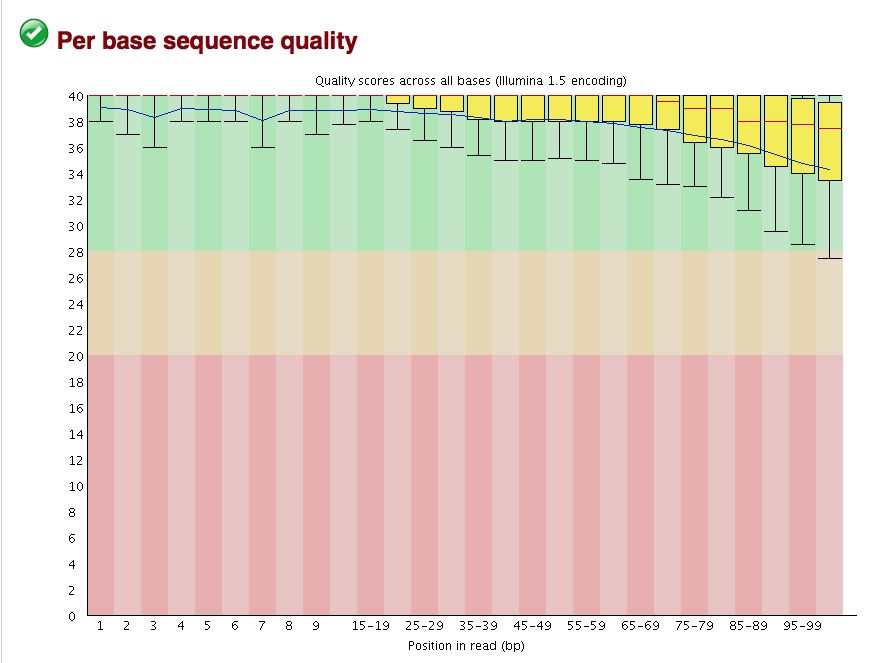
Take a look at the other areas that show up as quality issues. | |||
* FastQC flags potential problems | |||
* Are there any problems for us? | |||
[[File:CMV VAC WR 3 2 base quality.jpg]] | [[File:CMV VAC WR 3 2 base quality.jpg]] | ||
* It is imperative to understand your organism and its biology to interpret the data | |||
=== Quality Statistics and Boxplot === | |||
* | * Galaxy has its own set of tools for computing quality statistics (using R) | ||
* | * Generates raw statistics and a pretty box plot | ||
* Select NGS: QC and manipulation -> Compute Quality Statistics for CMV-21950R_3_1.fastq | |||
* Run it | |||
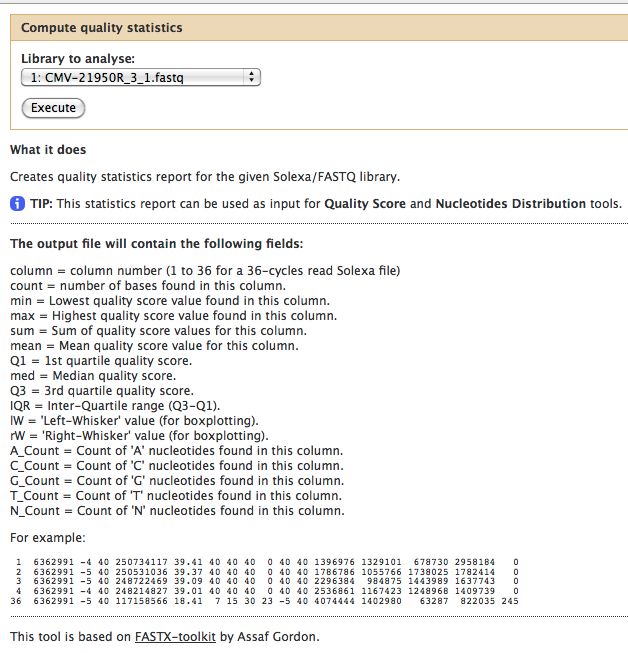
[[File:ComputeQualityStatsCMV21950R 3 1.jpg]] | |||
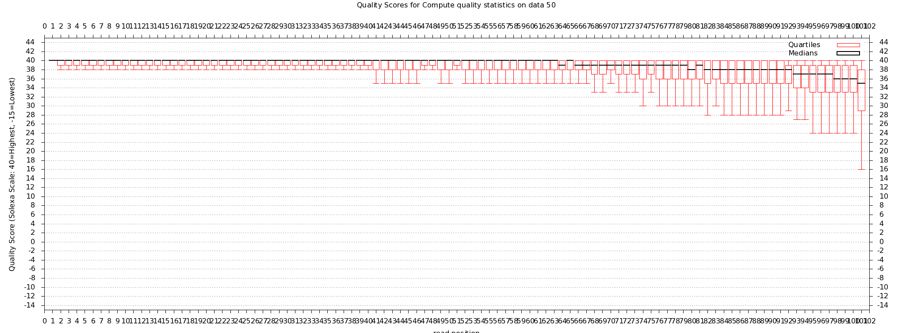
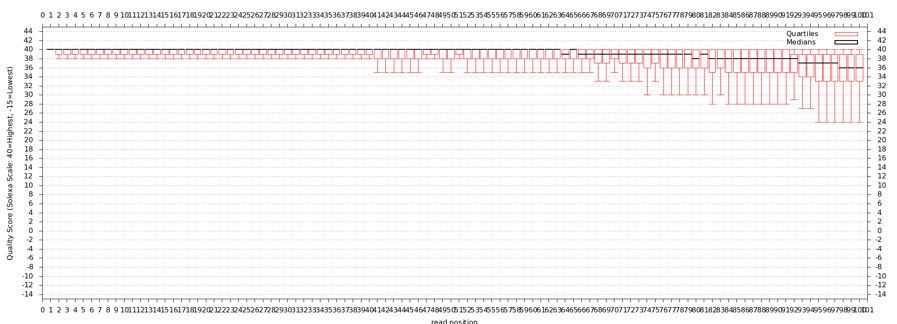
* Now draw a box and whiskers plot | |||
* "NGS: QC and manipulation" -> Draw Quality Score Boxplot | * "NGS: QC and manipulation" -> Draw Quality Score Boxplot | ||
[[File:DrawQualityScoreBoxplot.jpg]] | [[File:DrawQualityScoreBoxplot.jpg]] | ||
Revision as of 19:04, 15 September 2011
Galaxy DNA-Seq Tutorial
Linking to data
Link in the Mark Pritchard Vaccinia virus data set.
- Start with a blank history, there should be no numbered items on the right hand side of the pane. Otherwise create a new history.
- Select "Shared Data" from the top of the screen to bring up the Shared Data screen
- Select Data Library
- Select "Pritchard Vaccinia WR FastQ Files" from the alphabetically sorted list
- Click on the subfolder top box to select all 6 files
- Select "import to current history"
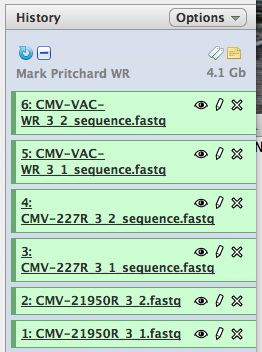
- Click on "Analyze Data" from the upper main menu. It should bring up the main page and your history pane should now look like the image below.
- Notice that with just 3 viruses are already over 4 GB of data!
Formatting and Grooming Data
- Your data is often NOT what you expect
FastQ Format
- FastQ format can mean many things, see the wikipedia entry on FastQ format
- Ilumina FastQ format also not informative, it changes
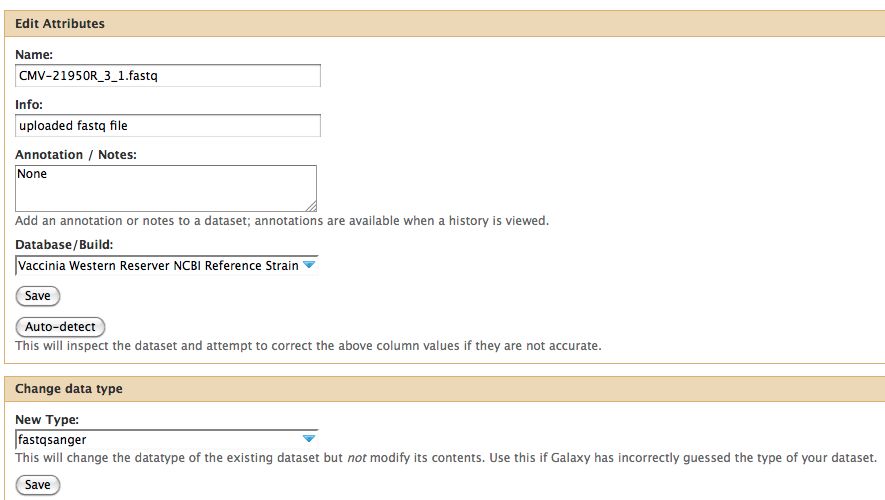
- Click on the pencil to icon in one of the virus images to pull up the attributes, your screen should look a bit like this:
- In Galaxy the expected data type of the galaxy tool must match EXACTLY with the data type in your history pane, otherwise the option to use that particular piece of data will not appear in the tool's drop down menu for data selection.
- Galaxy requires that everything go into Sanger format to be used. If you know your data is in sanger format, select fastqsanger for your data type. If it is not in that format, select fastq and run the FastQ Groomer.
- FastQ groomer is very useful, but slow
- Not a bad idea to run it if you are dealing with a new machine and have the time. Costly in terms of space.
Reference Genome
- Ensure the correct reference genome is selected
- WARNING - seeing the reference genome doesn't mean it is there for using
- The current list of genomes available at UAB is here
- Used for a variety of tools, it can be overridden earlier
- Any problems with CMV-21950R_3_1.fastq ?
Assessing the quality of the data
We will use a number of different tools from the "NGS: QC and manipulation" drop down menu. Try processing the FastQ files with:
- FastQC
- Compute Quality Statistics
FastQC
FastQC gives an attractive visual output and will flag potential problems.
- Select NGS: QC and manipulation -> FastQC
- Run it on all the fastq files
- Remember, you are on a cluster, this can be done in parrallel
Take a look at the other areas that show up as quality issues.
- FastQC flags potential problems
- Are there any problems for us?
- It is imperative to understand your organism and its biology to interpret the data
Quality Statistics and Boxplot
- Galaxy has its own set of tools for computing quality statistics (using R)
- Generates raw statistics and a pretty box plot
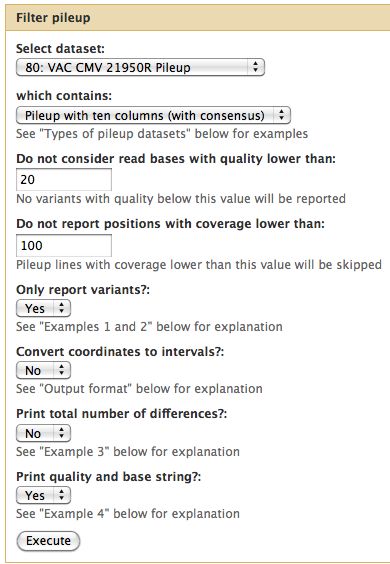
- Select NGS: QC and manipulation -> Compute Quality Statistics for CMV-21950R_3_1.fastq
- Run it
- Now draw a box and whiskers plot
- "NGS: QC and manipulation" -> Draw Quality Score Boxplot
Performing cleanup
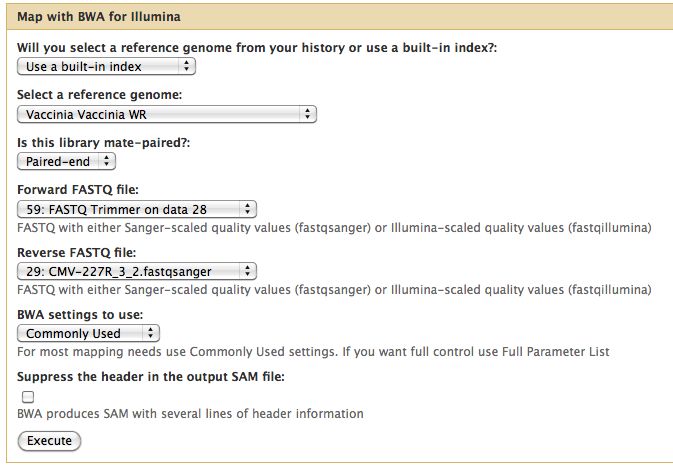
Short read alignment to reference genome using BWA
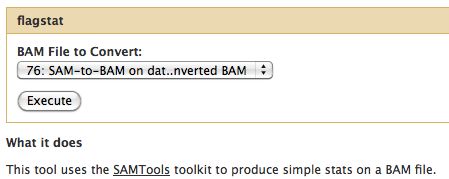
BAM Conversion
The output of many mapping programs is in SAM format and must be converted to the binary format. The conversion is lossless.
Variant Analysis
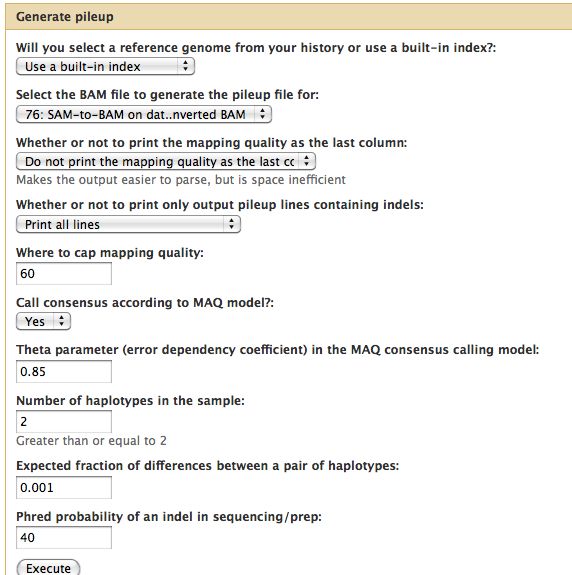
Samtools
- Essential toolset, performs a variety of functions including generation of a "pileup" file.
Other Variant Detection Approaches
- GATK - Emerging as the new best practise to call variants, not integrated into Galaxy just yet.
- SNPEff
- PATRIC, annotation sites
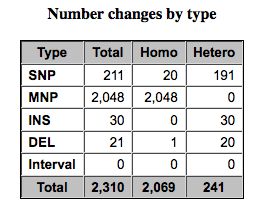
SNPEff and Variant Summation
Summarizing variants and effects of SNPs.
- Reference genome must be in SNPEff, need GFF3 file of your genome annotation
- SNPEff looks at heterozygous and homozygous SNPs, MNPs
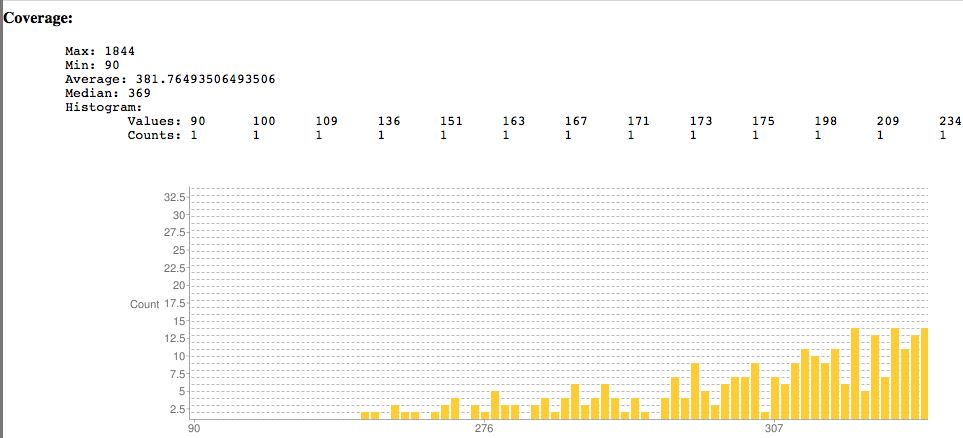
- Plots coverage, indel sizes
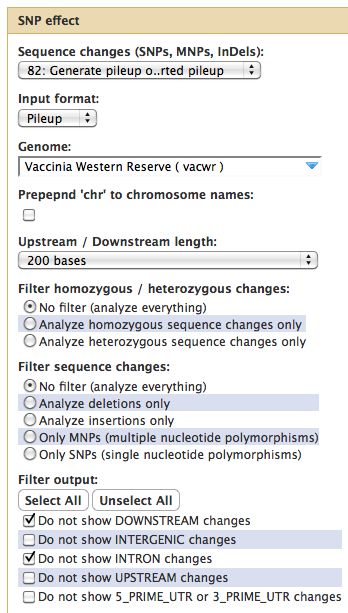
Running SNPEff
Results from SNPEff
- Only as good as your genome annotation file
- Introns in viruses?
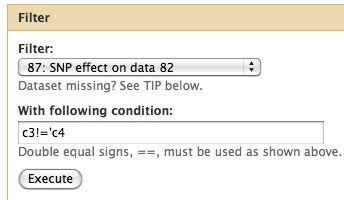
Locating Relevant SNPs versus Control